Diversifying the chloroquinoline scaffold against SARS-CoV-2 main protease: Virtual screening approach using cross-docking, SiteMap analysis and molecular dynamics simulation Scientific paper
Main Article Content
Abstract
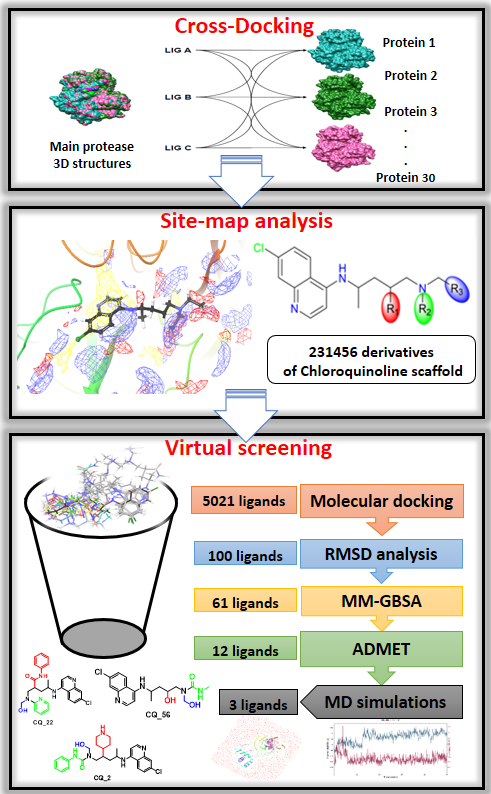
The absence of designated remedies for coronavirus disease 19 (Covid-19) and the lack of treatment protocols drove scientists to propose new small molecules and to attempt to repurpose existing drugs against various targets of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) in order to bring forward efficient solutions. The main protease (Mpro) is one of the most promising drug targets due to its crucial role in fighting viral replication. Several antiviral drugs have been used in an attempt to overcome the pandemic, such as hydroxychloroquine (HCQ). Despite its perceived positive outcomes in the beginning of the disease, HCQ was associated with several drawbacks, such as insolubility, toxicity, and cardiac adverse effects. Therefore, in the present study, a structure-based virtual screening approach was performed to identify structurally modified ligands of the chloroquinoline (CQ) scaffold with good solubility, absorption, and permeation aimed at eventually suggesting a more dependable alternative. PDB ID:7BRP Mpro was chosen as the most reliable receptor after cross-docking calculation using 30 crystal structures. Then, a SiteMap analysis was performed and a total of 231,456 structurally modified compounds of the CQ scaffold were suggested. After Lipinski criteria filtration, 64,312 molecules were docked and their MM-GBSA free binding energy were calculated. Next, ADME descriptors were calculated, and 12 molecules with ADME properties better than that of HCQ were identified. The resulting molecules were subjected to molecular dynamics (MD) simulation for 100 ns. The results of the study indicate that 3 molecules (CQ_22; CQ_2 and CQ_5) show better interactions and stability with the Mpro receptor. Binding interaction analysis indicates that GLU143, THR26, and HIS41 amino acids are potential binding hot-spot residues for the remaining 3 ligands.
Downloads
Metrics
Article Details

This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License.

Authors retain copyright and grant the journal right of first publication with the work simultaneously licensed under a Creative Commons Attribution license 4.0 that allows others to share the work with an acknowledgement of the work's authorship and initial publication in this journal.
Funding data
-
Direction Générale de la Recherche Scientifique et du Développement Technologique
-
Ministère de l'Enseignement Supérieur et de la Recherche Scientifique
Grant numbers Applied Organic Chemistry Laboratory (FNR 2000)
References
S. Ludwig,A. Zarbock, Anesth. Analg. 131 (1) (2020) 93 (https://doi.org/10.1213/ane.0000000000004845)
K. Dhama, S. K. Patel, K. Sharun, M. Pathak, R. Tiwari, M. I. Yatoo, Y. S. Malik, R. Sah, A. A. Rabaan,P.K. Panwar, Travel. Med. Infect. Dis. 37 (2020) 101830 (https://doi.org/10.1016/j.tmaid.2020.101830)
K. Yuki, M. Fujiogi,S. Koutsogiannaki, Clin. Immunol. 215 (2020) 108427 (https://doi.org/10.1016/j.clim.2020.108427)
I. M. Artika, A. K. Dewantari, A. Wiyatno, Heliyon. 6 (2020) e04743 (https://doi.org/10.1016/j.heliyon.2020.e04743)
Y. Zhou, Y. Hou, J. Shen, Y. Huang, W. Martin, F. Cheng, Cell Discov. 6 (2020) 14 (https://doi.org/10.1038/s41421-020-0153-3).
C. Wu, Y. Liu, Y. Yang, P. Zhang, W. Zhong, Y. Wang, Q. Wang, Y. Xu, M. Li, X. Li, M. Zheng, L. Chen,H. Li, Acta Pharm. Sin. B. 10 (2020) 766 (https://doi.org/10.1016/j.apsb.2020.02.008)
J. Shang, Y. Wan, C. Luo, G. Ye, Q. Geng, A. Auerbach,F. Li, Proc. Natl. Acad. Sci. U.S.A. 117 (2020) 11727 (https://doi.org/10.1073/pnas.2003138117).
L. Mousavizadeh,S. Ghasemi, J. Microbiol. Immunol. Infect. 54 (2021) 159 (https://doi.org/10.1016/j.jmii.2020.03.022)
J. Yang, S. J. L. Petitjean, M. Koehler, Q. Zhang, A. C. Dumitru, W. Chen, S. Derclaye, S. P. Vincent, P. Soumillion, D. Alsteens, Nat. Commun. 11 (2020) 4541 (https://doi.org/10.1038/s41467-020-18319-6)
X. Xue, H. Yu, H. Yang, F. Xue, Z. Wu, W. Shen, J. Li, Z. Zhou, Y. Ding, Q. Zhao, X.C. Zhang, M. Liao, M. Bartlam, Z. Rao, J. Virol. 82 (2008) 2515 (https://doi.org/10.1128/JVI.02114-07)
C. Liu, Q. Zhou, Y. Li, L.V. Garner, S.P. Watkins, L .J. Carter, J. Smoot, A. C. Gregg, A. D. Daniels, S. Jervey, D. Albaiu, ACS Cent. Sci. 6 (2020) 315 (https://doi.org/10.1021/acscentsci.0c00272)
M. T. ul Qamar, S. M. Alqahtani, M. A. Alamri,L.-L. Chen, J. Pharm. Anal. 10 (2020) 313 (https://doi.org/10.1016/j.jpha.2020.03.009)
Z. Jin, X. Du, Y. Xu, Y. Deng, M. Liu, Y. Zhao, B. Zhang, X. Li, L. Zhang, C. Peng, Y. Duan, J. Yu, L. Wang, K. Yang, F. Liu, R. Jiang, X. Yang, T. You, X. Liu, X. Yang, F. Bai, H. Liu, X. Liu, L.W. Guddat, W. Xu, G. Xiao, C. Qin, Z. Shi, H. Jiang, Z. Rao, H. Yang, Nature. 582 (2020) 289 (https://doi.org/10.1038/s41586-020-2223-y)
M. Bzówka, K. Mitusińska, A. Raczyńska, A. Samol, J. A. Tuszyński,A. Góra, Int. J. Mol. Sci. 21 (2020) 3099 (https://doi.org/10.3390/ijms21093099)
R. K. Harwansh, S. Bahadur, Curr. Pharm. Biotechnol. 23 (2022) 235 (https://doi.org/10.2174/1389201022666210322124348)
C. Scavone, S. Brusco, M. Bertini, L. Sportiello, C. Rafaniello, A. Zoccoli, L. Berrino, G. Racagni, F. Rossi, A. Capuano, Br. J. Pharmacol. 177 (2020) 4813 (https://doi.org/10.1111/bph.15072)
M. Nimgampalle, V. Devanathan, A. Saxena, J. Biomol. Struct. Dyn. 39 (2021) 4949 (https://doi.org/10.1080/07391102.2020.1782265)
P. Gautret, J. C. Lagier, P. Parola, V. T. Hoang, L. Meddeb, M. Mailhe, B. Doudier, J. Courjon, V. Giordanengo, V. E. Vieira, H. Tissot Dupont, S. Honoré, P. Colson, E. Chabrière, B. La Scola, J. M. Rolain, P. Brouqui, D. Raoult, Int. J. Antimicrob. Agents 56 (2020) 105949 (https://doi.org/10.1016/j.ijantimicag.2020.105949)
M. Wang, R. Cao, L. Zhang, X. Yang, J. Liu, M. Xu, Z. Shi, Z. Hu, W. Zhong, G. Xiao, Cell Res. 30 (2020) 269 (https://doi.org/10.1038/s41422-020-0282-0)
European Medicine Agency, COVID‐19: Chloroquine and hydroxychloroquine only to be used in clinical trials or emergency use programmes, 2020 (https://www.ema.europa.eu/en/news/covid-19-chloroquine-hydroxychloroquine-only-be-used-clinical-trials-emergency-use-programmes)
K. Sato, T. Mano, A. Iwata,T. Toda, Biosci. Trends 14 (2020) 139 (https://doi.org/10.5582/bst.2020.03082)
C. Chatre, F. Roubille, H. Vernhet, C. Jorgensen,Y.-M. Pers, Drug. Saf. 41 (2018) 919 (https://doi.org/10.1007/s40264-018-0689-4)
Z. Kashour, M. Riaz, M. A. Garbati, O. Al Dosary, H. Tlayjeh, D. Gerberi, M. H. Murad, M. R. Sohail, T. Kashour, I. M. Tleyjeh, J. Antimicrob. Chemother. 76 (2021) 30 (https://doi.org/10.1093/jac/dkaa403)
X. Cui, J. Sun, S.J. Minkove, Y. Li, D. Cooper, Z. Couse, P. Q. Eichacker, P. Torabi‐Parizi, Rev. Med. Virol. 31 (2021) e2228 (https://doi.org/10.1002/rmv.2228 )
T. Fiolet, A. Guihur, M. E. Rebeaud, M. Mulot, N. Peiffer-Smadja,Y. Mahamat-Saleh, Clin. Microbiol. Infect. 27 (2021) 19 (https://doi.org/10.1016/j.cmi.2020.08.022)
T. U. Singh, S. Parida, M. C. Lingaraju, M. Kesavan, D. Kumar, R. K. Singh, Pharmacol. Reports 72 (2020) 1479 (https://doi.org/10.1007/s43440-020-00155-6)
A. Khataniar, U. Pathak, S. Rajkhowa, A. N. Jha, Covid 2 (2022) 148 (https://doi.org/10.3390/covid2020011)
D. M. Teli, M. B. Shah, M. T. Chhabria, Front. Mol. Biosci. 7 (2021) 599079 (https://doi.org/10.3389/fmolb.2020.599079)
M. G. Santibáñez-Morán, E. López-López, F. D. Prieto-Martínez, N. Sánchez-Cruz, J. L. Medina-Franco, RSC Adv. 10 (2020) 25089 (https://doi.org/10.1039/D0RA04922K)
R. K. Gupta, E. L. Nwachuku, B. E. Zusman, R. M. Jha, A. M. Puccio, PLoS ONE. 16 (2021) e0257784 (https://doi.org/10.1371/journal.pone.0257784)
L. Fu, F. Ye, Y. Feng, F. Yu, Q. Wang, Y. Wu, C. Zhao, H. Sun, B. Huang, P. Niu, H. Song, Y. Shi, X. Li, W. Tan, J. Qi, G. F. Gao, Nat. Commun. 11 (2020) 4417 (https://doi.org/10.1038/s41467-020-18233-x)
E. F. Pettersen, T. D. Goddard, C. C. Huang, G. S. Couch, D. M. Greenblatt, E. C. Meng,T. E. Ferrin, J. Comput. Chem. 25 (2004) 1605 (https://doi.org/10.1002/jcc.20084)
H. M. Berman, J. Westbrook, Z. Feng, G. Gilliland, T. N. Bhat, H. Weissig, I. N. Shindyalov, P. E. Bourne, Nucleic Acids Res. 28 (2000) 235 (https://doi.org/10.1093/nar/28.1.235)
J. C. Shelley, A. Cholleti, L. L. Frye, J. R. Greenwood, M. R. Timlin, M. Uchimaya, J. Com¬put. Aided Mol. Des. 21 (2007) 681 (https://doi.org/10.1007/s10822-007-9133-z)
M. P. Jacobson, R. A. Friesner, Z. Xiang, B. Honig, J. Mol. Biol. 320 (2002) 597 (https://doi.org/10.1016/S0022-2836(02)00470-9)
T. A. Halgren, J. Chem. Inf. Model. 49 (2009) 377 (https://doi.org/10.1021/ci800324m)
T. A. Halgren, R. B. Murphy, R. A. Friesner, H. S. Beard, L. L. Frye, W. T. Pollard, J. L. Banks, J. Med. Chem. 47 (2004) 1750 (https://doi.org/10.1021/jm030644s).
M. F. Al Ajmi, M. T. Rehman, A. Hussain,G. M. Rather, Inter. J. Bio. Macromol. 116 (2018) 173 (https://doi.org/10.1016/j.ijbiomac.2018.05.023)
E. Harder, W. Damm, J. Maple, C. Wu, M. Reboul, J. Y. Xiang, L. Wang, D. Lupyan, M. K. Dahlgren, J. L. Knight, J. W. Kaus, D. S. Cerutti, G. Krilov, W. L. Jorgensen, R. Abel, R. A. Friesner, J. Chem. Theory Comput. 12 (2016) 281 (https://doi.org/10.1021/acs.jctc.5b00864)
A. O. Fadaka, R. T. Aruleba, N. R. S. Sibuyi, A. Klein, A. M. Madiehe, M. Meyer, J. Biomol. Struct. Dyn. 40 (2022) 3416 (https://doi.org/10.1080/07391102.2020.1847197)
K. J. Bowers, D. E. Chow, H. Xu, R. O. Dror, M. P. Eastwood, B. A. Gregersen, J. L. Klepeis, I. Kolossvary, M. A. Moraes, F. D. Sacerdoti, J. K. Salmon, Y. Shan, D. E. Shaw, in SC '06: Proceedings of the 2006 ACM/IEEE Conference on Supercomputing, Tampa, FL, 11–
–17 Nov. 2006, p. 43 (https://doi.org/10.1109/SC.2006.54).
L. Fu, F. Ye, Y. Feng, F. Yu, Q. Wang, Y. Wu, C. Zhao, H. Sun, B. Huang,P. Niu, Nat. Commun. 11 (2020) 1 (https://doi.org/10.1038/s41467-020-18233-x)
X. Yao, F. Ye, M. Zhang, C. Cui, B. Huang, P. Niu, X. Liu, L. Zhao, E. Dong, C. Song, S. Zhan, R. Lu, H. Li, W. Tan,D. Liu, Clin. Infect. Dis. 71 (2020) 732 (https://doi.org/10.1093/cid/ciaa237)
H. Rai, A. Barik, Y. P. Singh, A. Suresh, L. Singh, G. Singh, U. Y. Nayak, V. K. Dubey, G. Modi, Mol Divers. 25 (2021) 1905 (https://doi.org/10.1007/s11030-021-10188-5).