Application of liquid chromatography in defining the interaction of newly synthesized chalcones and related compounds with human serum albumin Scientific paper
Main Article Content
Abstract
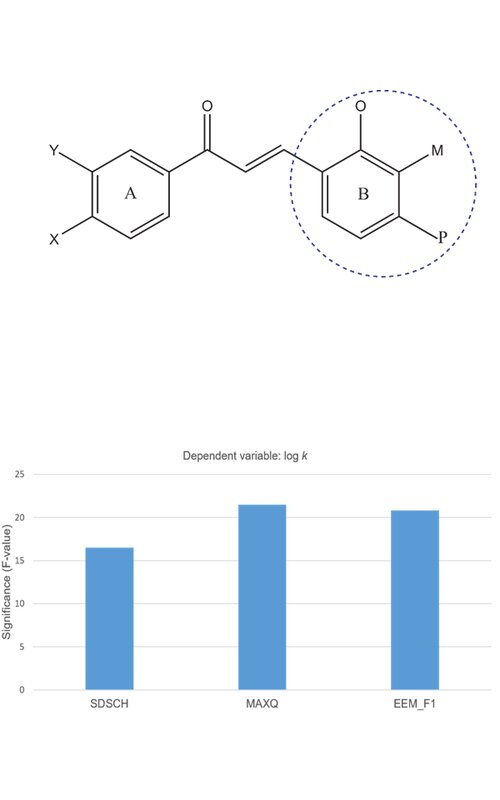
Defining the interaction of newly synthesized compounds with plasma proteins is an important step in the drug development process. Chromatographic techniques can be successfully used in predicting the biopharmaceutical and pharmacokinetic properties of newly synthesized compounds. The aim of this study is to investigate and isolate the most important molecular properties that affect the interaction of 20 newly synthesized chalcones and commercial compounds (lopinavir, ritonavir, darunavir and ivermectin) with human serum albumin (HSA). The retention behaviour of the selected compounds was tested on a CHIRALPAK®HSA column. A mixture of phosphate buffer (pH 7.0) and isopropanol (80:20 volume ratio) was used as the mobile phase, and the support vector method was used to form the quantitative structure retention relationship (QSRR) model. Based on the obtained values of retention parameters, it was observed that halogenated derivatives show the strongest, and methylated chalcone derivatives the weakest interaction with HSA. By correlating the retention and physicochemical properties of the tested compounds, it was shown that the structural (SDSCH) and electronic properties (MAXQ, EEM_F1) groups have the greatest influence on the retention behaviour and the interaction of the tested compounds with HSA. The obtained QSRR model can be applied in the prediction of the retention characteristics of new, structurally related chalcone derivatives on HSA stationary phase.
Downloads
Metrics
Article Details

This work is licensed under a Creative Commons Attribution 4.0 International License.

Authors retain copyright and grant the journal right of first publication with the work simultaneously licensed under a Creative Commons Attribution license 4.0 that allows others to share the work with an acknowledgement of the work's authorship and initial publication in this journal.
References
H. S. Kim, I. W. Wainer, J Chromatogr. B 870 (2008) 22 (https://doi.org/10.1016/j.jchromb.2008.05.029)
D. S. Hage, S. A. Tweed, J Chromatogr. B 699 (1997) 499 (https://doi.org/10.1016/S0378-4347(97)00178-3)
D. S. Hage, J. Anguizola, O. Barnaby, A. Jackson, M. J. Yoo, E. Papastavros, E. Pfaunmiller, M. Sobansky, Z. Tong, Curr. Drug Metab. 12 (2011) 313 (https://dx.doi.org/10.2174/138920011795202938)
J. Ghuman, P. A. Zunszain, I. Petitpas, A. A. Bhattacharya, M. Otagiri, S. Curry, J. Mol. Biol. 353 (2005) 38 (https://doi.org/10.1016/j.jmb.2005.07.075)
B. Ivkovic, PhD Thesis, University of Belgrade, Serbia, 2013 (http://doiserbia.nb.rs/phd/fulltext/BG20130320IVKOVIC.pdf)
Z. Nowakowska, Eur. J. Med. Chem. 42 (2007) 125 (https://doi.org/10.1016/j.ejmech.2006.09.019)
J. V. Basić, B. Ivković, S. Stevanović, A. Lazarević, Z. Vujić, Hem. Ind. 70 (2016) 511 (http://www.ache.org.rs/HI/2016/No5/HEMIND_Vol70_No5_p493-601_Sep-Oct_2016.pdf)
N. Turkovic, B. Ivkovic, J. Kotur-Stevuljevic, M. Tasic, B. Marković, Z. Vujic, Curr. Pharm. Des. 26 (2020) 802 (https://doi.org/10.2174/1381612826666200203125557)
Chem Axon, MarvinSketch, Version 6.1.0, Budapest, Hungary, 2013
Gaussian 09, Revision D.01, Gaussian, Inc., Wallingford CT, USA, 2009
ChemBio3D Ultra, Version 13.0, CambridgeSoft Corporation Cambridge, MA, USA, 2013
O. Trott, A. J. Olson, J Comput Chem. 31 (2010) 455 (https://doi.org/10.1002/jcc.21334)
SwissADME, http://www.swissadme.ch/index.php (accessed 15 Dec, 2021)
Statistica data analysis system, Version 13.6, Tulsa, OK, USA, 2019
A. Tropsha, Mol. Inform. 29 (2010) 476 (https://doi.org/10.1002/minf.201000061)
K. Roy, K. Supratik, A. Pravin, Chemometr. Intell. Lab. Syst. 145 (2015) 22 (https://doi.org/10.1016/j.chemolab.2015.04.013)
P. K. Ojha, I. Mitra, R. N. Das, Chemometr. Intell. Lab. Syst. 107 (2011) 194 (https://doi.org/10.1016/j.chemolab.2011.03.011)
DrugBank Online, https://go.drugbank.com/drugs/DB01601 (accessed 17 Feb, 2022)
DrugBank Online, https://go.drugbank.com/drugs/DB00503 (accessed 17 Feb, 2022)
DrugBank Online, https://go.drugbank.com/drugs/DB01264 (accessed 17 Feb, 2022)
D. Maksimovic-Ivanic, P. Fagone, J. McCubrey, K. Bendtzen, S. Mijatovic, F. Nicoletti, Int. J. Cancer. 140 (2017) 1713 (https://doi.org/10.1002/ijc.30529).