In silico identification of novel allosteric inhibitors of Dengue virus NS2B/NS3 serine protease Scientific paper
Main Article Content
Abstract
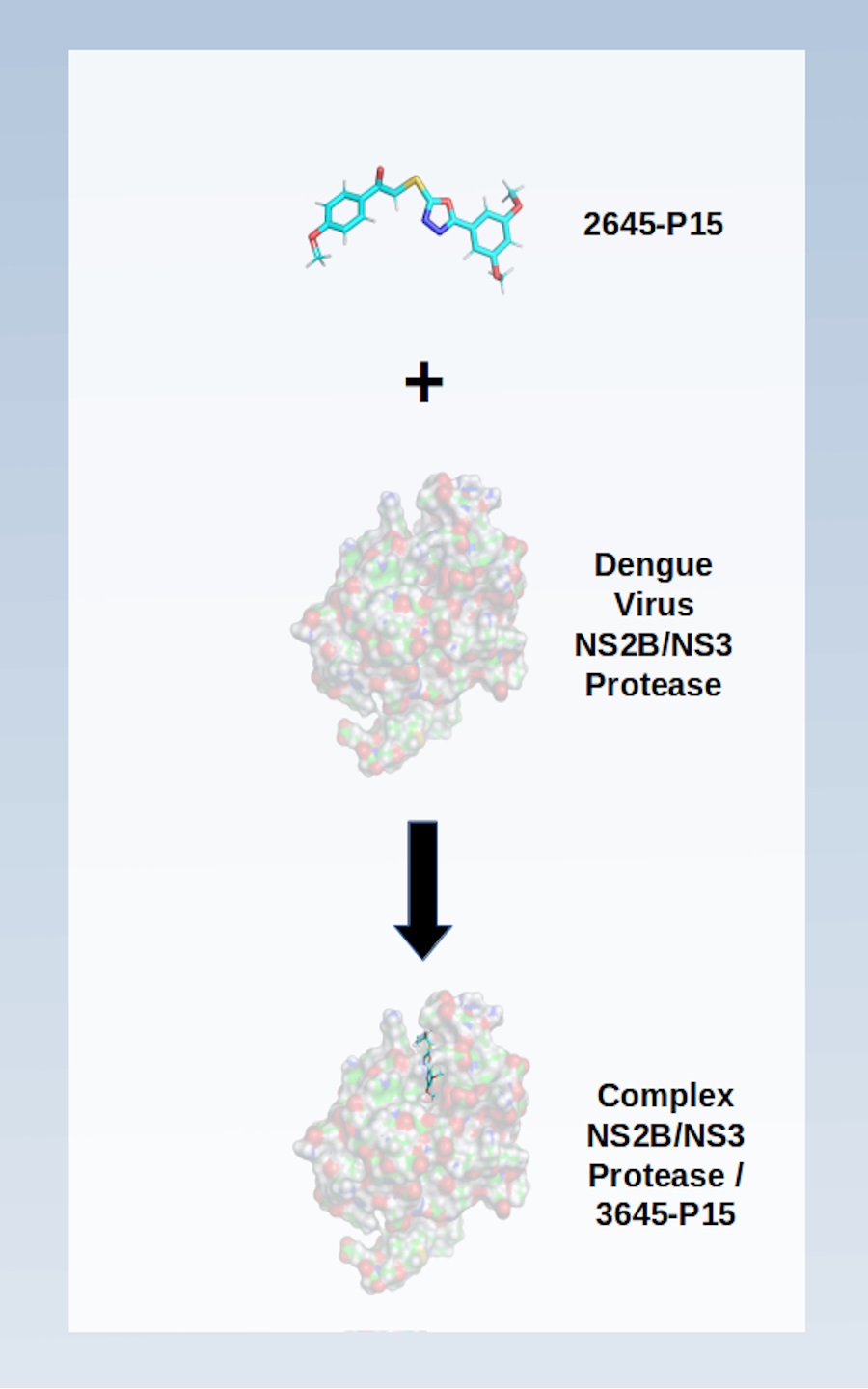
Although dengue is a disease that affects more than 100 countries and puts almost 400 million lives at risk each year, there is no approved antiviral in the treatment of this pathology. In this context, proteases are potential biological targets since they are essential in the replication process of this virus. In this study, a library of more than 3,000 structures was used to explore the allosteric inhibition of the NS2B/NS3 protease complex using consensual docking techniques. The results show four best ranked structures that were selected for molecular dynamics and free energy simulations. The present analysis corroborates with other studies (experimental and theoretical) presented in the literature. Thus, the computational approach used here proved to be useful for planning new inhibitors in the combat against Dengue disease.
Downloads
Metrics
Article Details

This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License.

Authors retain copyright and grant the journal right of first publication with the work simultaneously licensed under a Creative Commons Attribution license 4.0 that allows others to share the work with an acknowledgement of the work's authorship and initial publication in this journal.
References
A. Wilder-Smith, Murray, M. Quam, Clin. Epidemiol. (2013) 299 (https://doi.org/10.2147/CLEP.S34440)
M. G. Guzman E. Harris, Lancet 385 (2015) 453 (https://doi.org/10.1016/S0140-6736(14)60572-9)
S. Bhatt, P. W. Gething, O. J. Brady, J. P. Messina, A. W. Farlow, C. L. Moyes, J. M. Drake, J. S. Brownstein, A. G. Hoen, O. Sankoh, M. F. Myers, D. B. George, T. Jaenisch, G. R. W. Wint, C. P. Simmons, T. W. Scott, J. J. Farrar, S. I. Hay, Nature 496 (2013) 504 (https://doi.org/10.1038/nature12060)
C. P. Simmons, K. McPherson, N. Van Vinh Chau, D. T. Hoai Tam, P. Young, J. Mackenzie, B. Wills, Vaccine 33 (2015) 7061 (https://doi.org/10.1016/j.vaccine.2015.09.103)
K. V Pugachev, F. Guirakhoo, D. W. Trent, T. P. Monath, Int. J. Parasitol. 33 (2003) 567 (https://doi.org/10.1016/S0020-7519(03)00063-8)
D. Luo, S. G. Vasudevan, J. Lescar, Antiviral Res. 118 (2015) 148 (https://doi.org/10.1016/j.antiviral.2015.03.014)
M. Yildiz, S. Ghosh, J. A. Bell, W. Sherman, J. A. Hardy, ACS Chem. Biol. 8 (2013) 2744 (https://doi.org/10.1021/cb400612h)
B. Millies, F. von Hammerstein, A. Gellert, S. Hammerschmidt, F. Barthels, U. Göppel, M. Immerheiser, F. Elgner, N. Jung, M. Basic, C. Kersten, W. Kiefer, J. Bodem, E. Hildt, M. Windbergs, U. A. Hellmich, T. Schirmeister, J. Med. Chem. 62 (2019) 11359 (https://doi.org/10.1021/acs.jmedchem.9b01697)
M. Merdanovic, T. Mönig, M. Ehrmann, M. Kaiser, ACS Chem. Biol. 8 (2013) 19 (https://doi.org/10.1021/cb3005935)
M. Aminpour, C. Montemagno, J. A. Tuszynski, Molecules 24 (2019) 1693 (https://doi.org/10.3390/molecules24091693)
T. Casalini, J. Control. Rel. 332 (2021) 390 (https://doi.org/10.1016/j.jconrel.2021. 03.005)
B. S. Kolte, S. R. Londhe, B. R. Solanki, R. N. Gacche, R. J. Meshram, J. Mol. Graph. Model. 80 (2018) 95 (https://doi.org/10.1016/j.jmgm.2017.12.020)
P. Erbel, N. Schiering, A. D’Arcy, M. Renatus, M. Kroemer, S. P. Lim, Z. Yin, T. H. Keller, S. G. Vasudevan, U. Hommel, Nat. Struct. Mol. Biol. 13 (2006) 372 (https://doi.org/10.1038/nsmb1073)
R. Perez-Pineiro, A. Burgos, D. C. Jones, L. C. Andrew, H. Rodriguez, M. Suarez, A. H. Fairlamb, D. S. Wishart, J. Med. Chem. 52 (2009) 1670 (https://doi.org/10.1021/jm801306g)
M. D. de Oliveira, J. de O. Araújo, J. M. P. Galúcio, K. Santana, A. H. Lima, J. Mol. Graph. Model. 101 (2020) 107735 (https://doi.org/10.1016/j.jmgm.2020.107735)
E. Harigua-Souiai, Y. Z. Abdelkrim, I. Bassoumi-Jamoussi, O. Zakraoui, G. Bouvier, K. Essafi-Benkhadir, J. Banroques, N. Desdouits, H. Munier-Lehmann, M. Barhoumi, N. K. Tanner, M. Nilges, A. Blondel, I. Guizani, PLOS Negl. Trop. Dis. 12 (2018) e0006160 (https://doi.org/10.1371/journal.pntd.0006160)
G. Jones, P. Willett, R. C. Glen, J. Mol. Biol. 245 (1995) 43 (https://doi.org/10.1016/s0022-2836(95)80037-9)
S. R. Brozell, S. Mukherjee, T. E. Balius, D. R. Roe, D. A. Case, R. C. Rizzo, J. Comput. Aided Mol. Des. 26 (2012) 749 (https://doi.org/10.1007/s10822-012-9565-y)
M. Brecher, Z. Li, B. Liu, J. Zhang, C. A. Koetzner, A. Alifarag, S. A. Jones, Q. Lin, L. D. Kramer, H. Li, PLoS Pathog. 13 (2017) e1006411 (https://doi.org/10.1371/journal.ppat.1006411)
E. C. Meng, B. K. Shoichet, I. D. Kuntz, J. Comput. Chem. 13 (1992) 505 (https://doi.org/10.1002/jcc.540130412)
I. D. Kuntz, J. M. Blaney, S. J. Oatley, R. Langridge, T. E. Ferrin, J. Mol. Biol. 161 (1982) 269 (https://doi.org/10.1016/0022-2836(82)90153-X)
A. S. Pinheiro, J. B. C. Duarte, C. N. Alves, F. A. de Molfetta, Appl. Biochem. Biotechnol. 176 (2015) 1709 (https://doi.org/10.1007/s12010-015-1672-5)
J. M. Wang, R. M. Wolf, J. W. Caldwell, P. a Kollman, D. a Case, J. Comput. Chem. 25 (2004) 1157 (https://doi.org/10.1002/jcc.20035)
J. A. Maier, C. Martinez, K. Kasavajhala, L. Wickstrom, K. E. Hauser, C. Simmerling, J. Chem. Theory Comput. 11 (2015) 3696 (https://doi.org/10.1021/acs.jctc.5b00255)
P. A. Kollman, I. Massova, C. Reyes, B. Kuhn, S. Huo, L. Chong, M. Lee, T. Lee, Y. Duan, W. Wang, O. Donini, P. Cieplak, J. Srinivasan, D. A. Case, T. E. Cheatham, Acc. Chem. Res 33 (2000) 889 (https://doi.org/10.1021/ar000033j)
Gaussian 09 software, Pittsburgh, PA, 2016
R. Anandakrishnan, B. Aguilar, A. V. Onufriev, Nucleic Acids Res. 40 (2012) W537 (https://doi.org/10.1093/nar/gks375)
W. L. Jorgensen, J. Chandrasekhar, J. D. Madura, R. W. Impey, M. L. Klein, J. Chem. Phys. 79 (1983) 926 (https://doi.org/10.1063/1.445869)
T. Darden, D. York, L. Pedersen, J. Chem. Phys. 98 (1993) 10089 (https://doi.org/10.1063/1.464397)
J. P. Ryckaert, G. Ciccotti, H. J. C. Berendsen, J. Comput. Phys. 23 (1977) 327 (https://doi.org/10.1016/0021-9991(77)90098-5)
E. Wang, H. Sun, J. Wang, Z. Wang, H. Liu, J. Z. H. Zhang, T. Hou, Chem. Rev. 119 (2019) 9478 (https://doi.org/10.1021/acs.chemrev.9b00055)
P. A. Kollman, I. Massova, C. Reyes, B. Kuhn, S. Huo, L. Chong, M. Lee, T. Lee, Y. Duan, W. Wang, O. Donini, P. Cieplak, J. Srinivasan, D. A. Case, T. E. Cheatham, Acc. Chem. Res. 33 (2000) 889 (https://doi.org/10.1021/ar000033j)
S. Genheden U. Ryde, Expert Opin. Drug Discov. 10 (2015) 449 (https://doi.org/10.1517/17460441.2015.1032936)
R. A. Costa, J. N. Cruz, F. C. A. Nascimento, S. G. Silva, S. O. Silva, M. C. Martelli, S. M. L. Carvalho, C. B. R. Santos, A. M. J. C. Neto, D. S. B. Brasil, Med. Chem. Res. 28 (2019) 246 (https://doi.org/10.1007/s00044-018-2280-z)
E. P. Semighini, J. A. Resende, P. de Andrade, P. A. B. Morais, I. Carvalho, C. A. Taft, C. H. T. P. Silva, J. Biomol. Struct. Dyn. 28 (2011) 787 (https://doi.org/10.1080/07391102.2011.10508606)
M. Hariono, S. B. Choi, R. F. Roslim, M. S. Nawi, M. L. Tan, E. E. Kamarulzaman, N. Mohamed, R. Yusof, S. Othman, N. Abd Rahman, R. Othman, H. A. Wahab, PLOS One 14 (2019) e0210869 (https://doi.org/10.1371/journal.pone.0210869)
A. J. Fathima, G. Murugaboopathi, P. Selvam, Curr. Bioinform. 13 (2018) 606 (https://doi.org/10.2174/1574893613666180118105659)
A. Mukhametov, E. I. Newhouse, N. A. Aziz, J. A. Saito, M. Alam, J. Mol. Graph. Model. 52 (2014) 103 (https://doi.org/10.1016/j.jmgm.2014.06.008)
F. A. D. M. Opo, M. M. Rahman, F. Ahammad, I. Ahmed, M. A. Bhuiyan, A. M. Asiri, Sci. Rep. 11 (2021) 4049 (https://doi.org/10.1038/s41598-021-83626-x)
A. Fornili, F. Autore, N. Chakroun, P. Martinez, F. Fraternali, Computational Drug Discovery and Design, Springer, New York, 2011, p. 375 (https://doi.org/10.1007/978-1-61779-465-0_23)
M. Shahbaaz, A. Nkaule, A. Christoffels, Sci. Rep. 9 (2019) 4405 (https://doi.org/10.1038/s41598-019-40621-7)
F. Ghasemi, A. Zomorodipour, A. A. Karkhane, M. R. Khorramizadeh, J. Mol. Graph. Model 68 (2016) 39 (https://doi.org/10.1016/j.jmgm.2016.05.011)
H. Wu, S. Bock, M. Snitko, T. Berger, T. Weidner, S. Holloway, M. Kanitz, W. E. Diederich, H. Steuber, C. Walter, D. Hofmann, B. Weißbrich, R. Spannaus, E. G. Acosta, R. Bartenschlager, B. Engels, T. Schirmeister, J. Bodem, Antimicrob. Agents Chemother. 59 (2015) 1100 (https://doi.org/10.1128/AAC.03543-14)
R. A. Costa, J. N. Cruz, F. C. A. Nascimento, S. G. Silva, S. O. Silva, M. C. Martelli, S. M. L. Carvalho, C. B. R. Santos, A. M. J. C. Neto, D. S. B. Brasil, Med. Chem. Res. 28 (2019) 246 (https://doi.org/10.1007/s00044-018-2280-z)
B. Millies, F. von Hammerstein, A. Gellert, S. Hammerschmidt, F. Barthels, U. Göppel, M. Immerheiser, F. Elgner, N. Jung, M. Basic, C. Kersten, W. Kiefer, J. Bodem, E. Hildt, M. Windbergs, U. A. Hellmich, T. Schirmeister, J. Med. Chem. 62 (2019) 11359 (https://doi.org/10.1021/acs.jmedchem.9b01697)
R. Othman, T. S. Kiat, N. Khalid, R. Yusof, E. Irene Newhouse, J. S. Newhouse, M. Alam, N. A. Rahman, J. Chem. Inf. Model. 48 (2008) 1582 (https://doi.org/10.1021/ci700388k)
D. Popovic I. Djordjevic, J. Serb. Chem. Soc. 85 (2020) 1429 (https://doi.org/10.2298/JSC200720047P)
D. M. Popović A. A. Stuchebrukhov, J. Am. Chem. Soc. 126 (2004) 1858 (https://doi.org/10.1021/ja038267w).